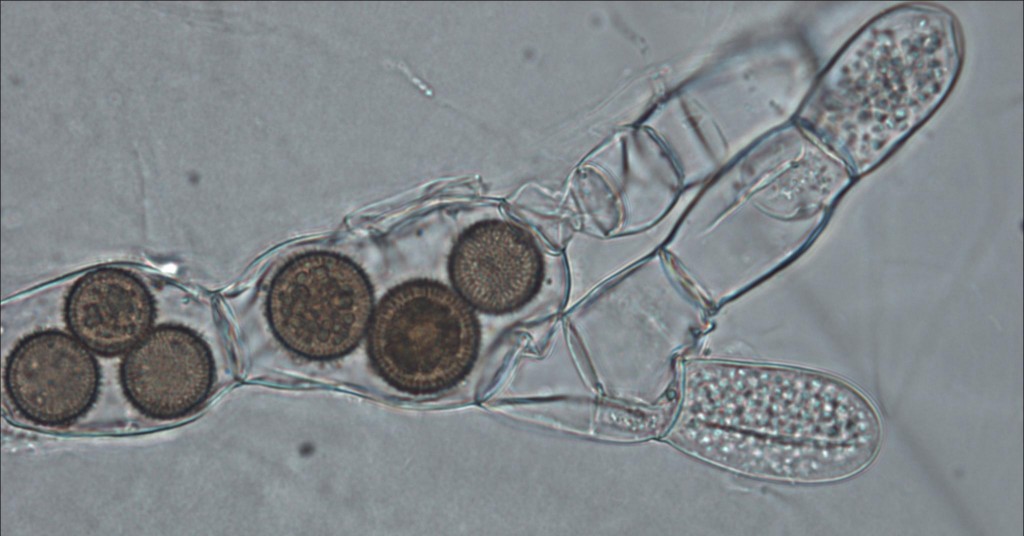
Rozella sp., a previously discovered internally parasitic cryptomycete totally devoid of a chitin cell wall. I think they're the brown truffley balls. Click to enlarge. Creative Commons Timothy Y. James. License at http://tolweb.org/Rozella_spp./103935
To listen to news reports, you might have thought the big news was that the fungi have an interesting new set of oddball relatives. But these news reports completely missed what I consider the most earth-shattering news in the paper in Nature announcing the discovery, cleverly hidden at the end of this sentence:
Our analyses also recovered a highly diverse clade of environmental sequences branching with the fungi and demonstrated that current models of fungal evolution and biodiversity, which are largely based on cultured microbes, have missed a huge fraction of the kingdom (perhaps even approaching half).
Wait– hold up. HALF?!! Thus, the news isn’t that there’s some new oddball fungal relatives out there. It’s that the new oddball fungal relatives may make up half of Kingdom Fungi. And the first sample of this huge, undiscovered and massively diverse group came from none other than one of the scientists’ very own university pond. In spite of our best efforts to survey life, we had virtually no idea these things were out there — in virtually every environment we have now looked.
To understand how we may have been ignorant of half of Kingdom Fungi, you must think back to the time before DNA. To discover new life that couldn’t be seen with a hand lens, you had two options:
- Take some dirt or water. Wash and dump said sample onto a microscope slide. Apply various dyes as desired. Squash under coverslip. Place under microscope and stare intensely, searching for something that might be alive. Repeat thousands of times. See doctor about eyestrain.
- Take some dirt or water. Dump said sample into a petri dish containing some food-like substance. Wait. Pray the smell is not too bad. Place sample of stuff that grows on microscope slide. Squash under coverslip. Place under microscope and stare at similar-looking blobs. Repeat thousands of times. See doctor about eyestrain.
As you can see, this system left a lot to be desired. And it’s how hundreds of groups of organisms got mis-classified with other groups, because all biologists had to go on was the way the microbes looked and — in some cases, behaved — under a microscope. In addition, many groups of life on Earth are not culturable in a petri dish, or have evolved to look and behave in very similar ways, regardless of their ancestry.
Enter DNA. With this molecule, we can detect true heredity by comparing how the unique sequence of four letters in the code have gradually changed over time in related organisms. With millions of positions for letters in a defined order (like a cryptokey or license plate), it is easy to tell who is related to who and what is unique. By doing massive sampling of all the DNA in the environment, we can now easily sample for and pick out new variations of known genes (and the new organisms they must represent) in dirt and water samples, because they will look unique even if the organisms that make them do not*. And we can compare those new sequences** to sequences of organisms whose appearances and taxonomies we understand — in this case, a broad selection of fungi and their closest relatives.
AND we can use sequences unique to new groups as organismal metal detectors (scientists call them “probes”) to finger them under the microscope so we can see what they actually look like. Since DNA is a pair of complementary strands that bind to each other, you can make a probe that’s complementary to the particular sequence you’re looking for*** and attach that bit of DNA to a fluorescent molecule. Then you stick it in your hopelessly confusing sample of water or dirt and turn out the lights. Voila! Your organism lights up like a Christmas tree! No more eyestrain.
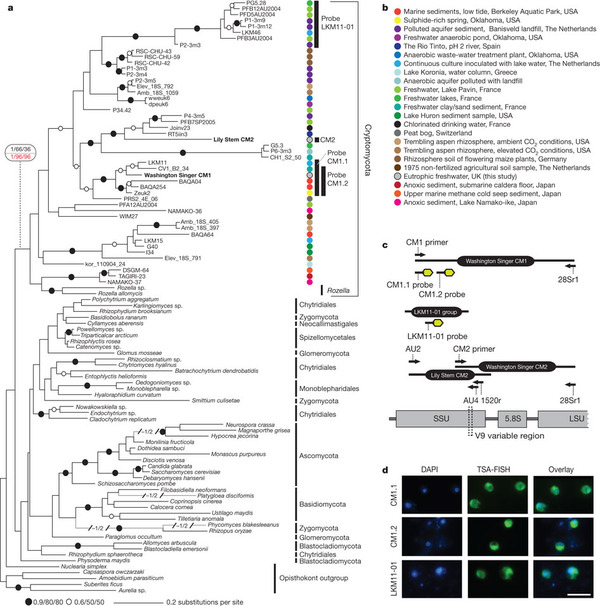
And that is just what a group of enterprising mycologists did. Biologists have sampled many different environments for DNA from all sources. In this study, scientists accessed online records of those sequences and looked for any that seemed to fit in with the fungi. They were in for a shock. As it turned out, there was an enormous group of organisms that branched with the fungi, that apparently no one had ever seen or cultured before, with the exception of one genus. Here is what they found:
This figure is actually much simpler than it looks. All the branches at the top with the colored dots represent organisms new to science in the new group, dubbed the “Cryptomycota”, or “hidden fungi”. There is one exception: a group of strange parasites called Rozella, which was already known. Everything else with the exception of the “opisthokont outgroup” — the group most closely related to Cryptomycetes and Fungi — are organisms from the known Kingdom Fungi. The rainbow of colors give you a feel for all the environments from which they found this new group. The only place they did not find cryptomycetes was seawater, although they did find them in mud on the seabed. The branch lengths should give you a feel for the diversity. It is incredible. The breadth of ribosomal DNA molecular diversity shown in they cryptomycetes here is similar to that of the known fungal kingdom.
What did these critters look like once fluorescently tagged by DNA? So far, the cells found in samples were 3-5 micrometers long — not much bigger than bacteria — and capable of making a microtubule-based flagellum like other good eukaryotes. In two of the three groups of cyrptomycetes they studied with probes, they found more than half of the cells had single flagella. Other, non-flagellate examples seemed to like latching onto — of all things — diatoms, the algae that make glass pillbox houses in a blinding array of uber-cute shapes. In addition, though they clearly group with the fungi, the Cryptomyctes — including Rozella — lack a very important marker of fungus-hood: they have no chitin/cellulose-rich cell wall.
For fungi as we know it, this is about as elemental as it gets. Their distinctive tubular, chitinous cell walls have enabled them to infiltrate a variety of foodstuffs that other creatures cannot and digest them osmotically. That is, grow into food that is concentrated in sugars or other nutrients that would make other cells shrivel up and die thanks to osmosis if they attempted it. We humans exploit this effect to preserve our food. Pickles, preserves, and peanut butter are so rich in salt, sugar, and general osmotic potential respectively that no bacteria or fungi can survive. But in nature, there’s a gray area where only fungi are bold enough to tread. They grow into their food (i.e. log, leaf, leftovers, etc.) They excrete digestive enzymes. They absorb the resulting goo. Life is good.
But the cryptomycetes have no such chitinous cell wall. The researchers know this because they stained the cells that probes identified as cryptomycetes with chemicals that bind to chitin and cellulose, the stuff of fungal cell walls, and they lit up like a Christmas tree during a power outage. Which is to say, they did not.
This is a problem for our definition of Fungi. As I learned it, to be a fungus requires five things:
- you are eukaryotic (you have a nucleus where you keep your DNA and various other cute organelles orbiting it)
- you are heterotrophic (you get food by taking it, rather than making it)
- you are absorptive (we already covered this)
- you reproduce with spores (a much better method than that used by mammals, IMO)
- you possess a tubular body with a chitin cell wall
Yeast are somewhat exceptions to these last two; they are spherical and reproduce by budding. But they do not make up the majority of the fungi, by far, and so far, no one has ever bucked the chitin rule.
Hmmmm.
This brings us to the real questions wrought by this study: Are the Cryptomycota truly fungi? Or are they a larger group to which the fungi belong? Or are they a sister group of fungi? Did they ever have chitin cell walls or have they lost them? Are there vestiges of chitin in ephemeral life stages the researchers may have missed? There has already been some commentary on these questions in the comments here. We may learn a lot about the evolution of fungi based on the answers. One thing seems clear: if we decide they are “Fungi”, our definition — and our perceptions of the group — must change.
Aside from these important questions, we also have the delicious prospect of discovering how the contents of this new clade — the Cryptomycota — look and behave. There is almost no telling what we might find among some of the more eccentric members, though based on their fluorescence experiments so far, the paper’s authors suggest a saprophytic or parasitic lifestyle for the majority of species based on what they’ve actually seen under the microscope: a free swimming zoospore stage with a single tail, a resting cyst stage, and period of attachment and feasting upon a “second-party cell”, a.k.a. a helpless diatomic host or rotting microbial corpse (like leeches — or hyenas). But the group is so genetically diverse there are probably many, many ways of life. Based on what we know of the fungi, the authors suggest, there may be many life stages and accessories yet to discover: germ tubes, root-like food-harvesting devices called rhizoids, and spore-making houses called sporangia, filter-feeding structures, DirectTV dishes, etc.
Without a cell wall, they may even be phagocytic — that is, able to engulf, swallow, and digest prey internally. Outside the Cryptomycota, fungi’s closest sister group is the nucleariids, a group of amoebae with filamentous (hmmm . . . ) pseudopods that feed, as amoebae do, by phagocytosis. Makes a girl think.
As the biologists say, this is an exciting time to study life. When reports like this come along, I feel a bit like a mapmaker in 16th Century Portugal hungrily studying the blank spots on the map. Can’t. Wait.
_________________________________________________________________
* because those groups will have two very different evolutionary histories, and thus, genetic patterns.
** in this case, of ribosomal RNA genes, a commonly sampled subject because they change so slowly and are present in everything
*** in this study, they used ten 18-base pair long clade-specific probes for taxonomic analysis fluorescent tagging purposes
![]()
Jones MD, Forn I, Gadelha C, Egan MJ, Bass D, Massana R, & Richards TA (2011). Discovery of novel intermediate forms redefines the fungal tree of life. Nature PMID: 21562490

{ 1 trackback }
{ 6 comments… read them below or add one }
Wonderful news — things like this keep me eagerly reading your blog.
I just spent some time searching the web for other stories about this, and most of them are pretty poor, even dragging out the term “missing link”; but there’s one more good blog post at Southern Fried Science. Plus, the German news article at Spektrum der Wissenschaft is rather better than most of the English-language ones.
I am going to be lazy and not RTFA. How likely is it that these cryptomycetes are multicellular? If not, is it likely that the organisms in this group are the unicellular closest relatives to fungi, in much the same way that choanoflagellates are/may be the closest unicellular relatives to animals? That would solve the nomenclature issues and return us to the status quo of there are a lot of more-or-less-unicellular heterotrophs that do weird stuff . It still opens up exciting possibilities about understanding one of the origins of multicellularity. Where do slime ‘molds’ fit in in this phylogeny?
So far the the ones they’ve probed have all stained unicellular. But their probes only sampled a small fraction of the group, so who knows what the rest are like. To answer your second question, yes. Many of the most basal fungi — the chytrids — are unicellular for most or all of their life too. Of course, they have chitinous cell walls.
Slime molds, as usual, are off in left field in the Amoebozoa; see here for one conservative tree.
Sorry for the delay in responding — just got back today from my talk in the Springs!
The same has happened in the viral world. We have missed many viruses because they cannot be cultured. Try to plate sea water on E. coli and you won’t get much in the way of plaques; sequence the water, and you find one million phages per ml. The same goes for rhinoviruses, agents of the common cold. We knew of 100 culturable rhinoviruses until sequencing of snot revealed another 150. And to think I spend one year sequencing a 7.5 kb viral genome in 1981!
SO FANTASTIC. i don’t know what else to do but use all caps. :) makes me wonder about so many things! and just like vasha, this is the kind of topic (and lovely writing) that makes your blog one of my absolute favorites. and i tell so many people about it. mostly i get a lot of funny looks wondering why this mom of two has a “favorite fungi/biodiversity blog” -ha!
and vincent, you bring up so many good points, too! reading all this, i pretty much just assume that not finding anything in the seawater just means we haven’t found whatever it is we don’t know to look for yet. this planet is crawling with fun stuff! if i could turn my bonus room into a serious amateur lab, i totally would! haha!